 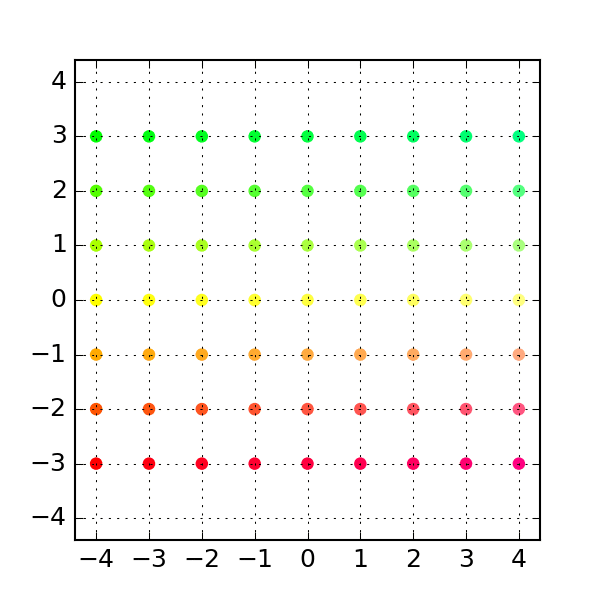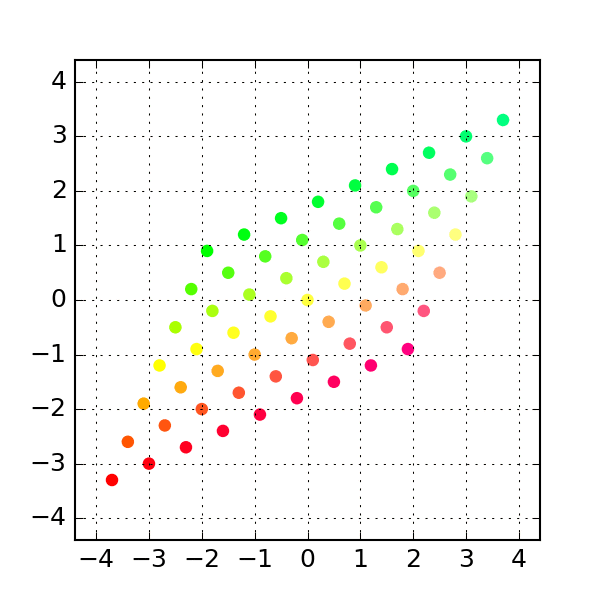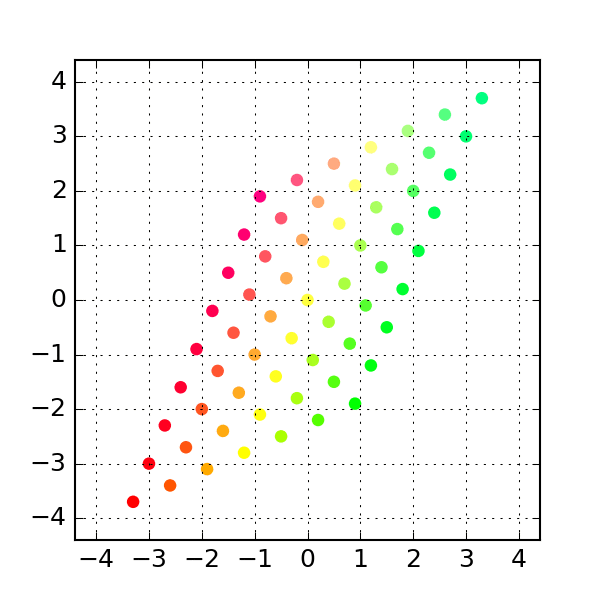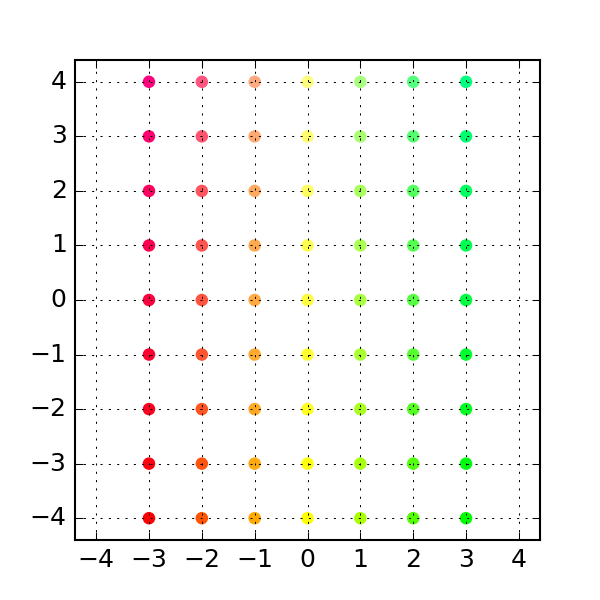
'''Generates indices, making random stratified split into training set and testing sets import numpy as npĭef get_train_test_inds(y,train_proportion=0.7): This makes training and testing sets better reflect the properties of the original dataset. Startified division also generates training and testing set randomly but in such a way that original class proportions are preserved. You may also consider stratified division into training and testing set. sklearn also includes more advanced "stratified sampling" methods that create a partition of the data that is balanced with respect to some features, for example to make sure that there is the same proportion of positive and negative examples in the training and test set. Many of those are available in the sklearn library (k-fold, leave-n-out. There are many ways other ways to repeatedly partition the same data set for cross validation. Training_idx, test_idx = indices, indices If speed becomes an issue, I would go with the Cython solution.If you want to split the data set once in two parts, you can use, or if you need to keep track of the indices (remember to fix the random seed to make everything reproducible): import numpy In my opinion, it is less obscure than argsort, and also faster for large input sizes. def invert_permutation(p):ġ00000 loops, best of 3: 11.6 µs per loopġ00000 loops, best of 3: 11.5 µs per loopĪll in all, I would go with the Short answer approach mentioned at the top for code clarity. I tested it with Python 3.5 and NumPy 1.11 on the machine that I was using back in 2014. Jamie says it was already resolved in NumPy 1.9. Jamie, Andris and Paul pointed out in comments below that the performance issue with fancy indexing was resolved. So, the np.put solution is still not as fast as possible (ran 12.8 ms for this input size argsort took 72.7 ms). 
This is most likely because we allocate and fill in an extra array (at the np.arange() call) with the np_put approach.Īlthough you didn't ask for a Cython solution, just out of curiosity, I also timed the following Cython solution with typed memoryviews: import numpy as np 
To be fair, np.argsort still beats the np.put approach for smaller n (the tipping point is around n = 1210 on my machine): p = create_input(1210) This is a nice 5.6x speed up for next to nothing! Which gives for n = 700 000 (the same size as above): p = create_input(700000) However, there is a less straightforward way to vectorize the above for loop with np.put: def np_put(p): O(n) for the single-pass algorithm) and the single-pass algorithm will be consistently faster after a sufficiently large n = p.size (threshold is around 700k on my machine). S = np.zeros(p.size, p.dtype) # np.zeros is better than np.empty here, at least on Linuxįrom my IPython notebook: p = create_input(700000)Įventually the asymptotic complexity kicks in ( O(n log n) for argsort vs. An update with NumPy 1.11 follows later.)Ī single-pass, linear time algorithm is expected to be faster than np.argsort interestingly, the trivial vectorization ( s = xrange(p.size), see index arrays) of the above for loop is actually slightly slower than np.argsort as long as p.size < 700 000 (well, on my machine, your mileage will vary): import numpy as np (The original answer from the timings are valid for NumPy 1.8. If you just want to know the conclusion, jump to the end of this answer. The rest of the answer is concerned with the efficient vectorization of the above for loop. This is just a single-pass, linear time algorithm with constant memory requirement: from _future_ import print_function P = np.asanyarray(p) # in case p is a tuple, etc. The array_like argument p must be some permutation of 0, 1. """Return an array s with which np.array_equal(arr, arr) is True. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
